Author: Abhishek Das, North Carolina State University
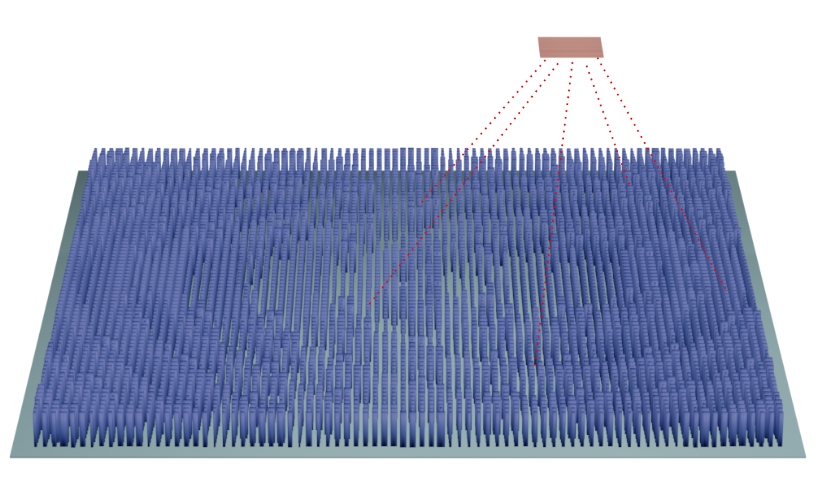
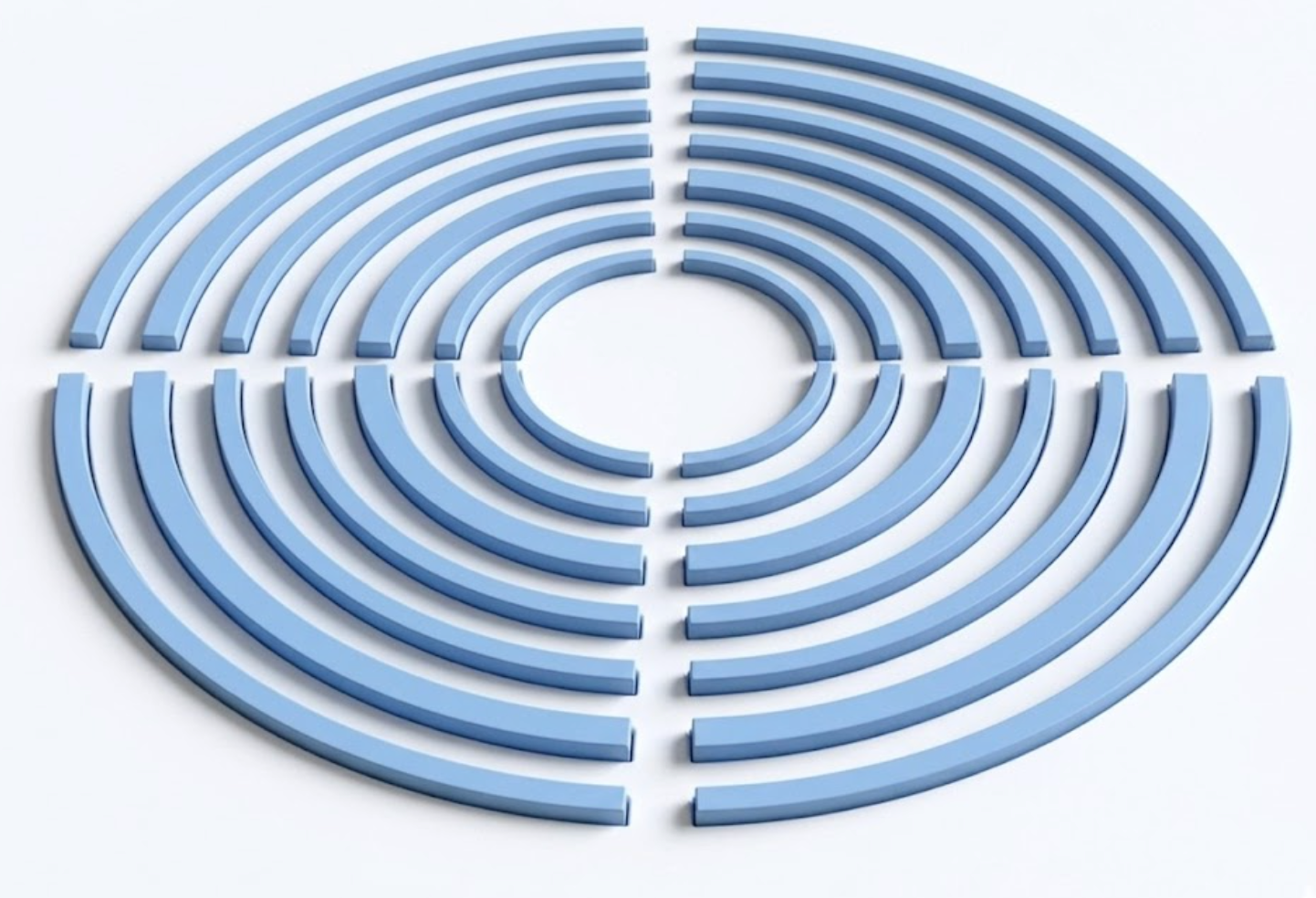
In this notebook, we simulate a bullseye grating on a GaAs slab using Tidy3D. A point dipole source excites the structure, and the far-field radiation pattern is computed to evaluate Gaussian mode overlap and power collection efficiency within a target numerical aperture. Resonance analysis is also performed to identify the high-Q modes of the structure.
The bullseye geometry is defined using GDS (gdstk) and imported into Tidy3D.

import numpy as np
import matplotlib.pyplot as plt
import tidy3d as td
import tidy3d.web as web
from tidy3d.plugins.resonance import ResonanceFinder
import gdstk
Set Up the Simulation¶
lambda_min = 0.8
lambda_max = 1.1
monitor_lambda = np.linspace(lambda_min, lambda_max, 401)
monitor_freq = td.constants.C_0 / monitor_lambda
automesh_per_wvl = 40
sim_size = Lx, Ly, Lz = (5.5, 5.5, 2)
offset_monitor = 0.4
vacuum = td.Medium(permittivity = 1, name = 'vacuum')
GaAs_permittivity = 3.55**2
GaAs = td.Medium(permittivity = GaAs_permittivity, name = 'GaAs')
freq0 = 302.675e12
fwidth = 144.131e12
dipole_source = td.PointDipole(
center = (0, 0, 0),
source_time = td.GaussianPulse(
freq0 = freq0,
fwidth = fwidth
),
polarization = 'Ex',
name = 'dipole_source'
)
t_start = 10/fwidth
t_stop = 30e-12
Define Structures¶
slab_side_length = 5
slab_height = 0.2
slab = td.Structure(
geometry = td.Box(
center = (0, 0, 0),
size = (slab_side_length, slab_side_length, slab_height),
),
medium = GaAs
)
lib = gdstk.Library()
cell = lib.new_cell("BULLSEYE")
central_inner_radius = 0.355 + 0.014
ring_width = 0.07 + np.array([-0.036, -0.015, +0.011, -0.009, -0.011, +0.012, +0.005])
ring_height = 0.2
ring_period = 0.1075
angle = 3
quadrants = 4
theta1 = angle * np.pi / 180
theta2 = (90 - angle) * np.pi / 180
ring_numbers = np.size(ring_width)
x_origin = 0
y_origin = 0
inner_radius = central_inner_radius
for i in range(ring_numbers):
outer_radius = inner_radius + ring_width[i]
for j in range(quadrants):
gc_slice = gdstk.ellipse(
(x_origin, y_origin),
inner_radius,
outer_radius,
theta1 + j*np.pi/2,
theta2 + j*np.pi/2,
layer = 1,
datatype = 1,
tolerance = 0.0001
)
cell.add(gc_slice)
inner_radius = outer_radius + ring_period
gc_etch = td.Geometry.from_gds(
cell,
gds_layer = 1,
axis = 2,
slab_bounds = (-ring_height/2, ring_height/2)
)
etch = td.Structure(
geometry = gc_etch,
medium = vacuum,
name = "etch"
)
Define Monitor¶
time_monitor_1 = td.FieldTimeMonitor(
name = "time_monitor_1",
size = [0.0, 0.0, 0.0],
center = [0.1, 0.1, 0.0],
start = t_start
)
time_monitor_2 = td.FieldTimeMonitor(
name = "time_monitor_2",
size = [0.0, 0.0, 0.0],
center = [0.1, 0.2, 0.0],
start = t_start
)
time_monitor_3 = td.FieldTimeMonitor(
name = "time_monitor_3",
size = [0.0, 0.0, 0.0],
center = [0.2, 0.2, 0.0],
start = t_start
)
time_monitor_4 = td.FieldTimeMonitor(
name = "time_monitor_4",
size = [0.0, 0.0, 0.0],
center = [0.2, 0.1, 0.0],
start = t_start
)
time_monitors = [time_monitor_1, time_monitor_2, time_monitor_3, time_monitor_4]
far_field_monitor = td.FieldProjectionAngleMonitor(
center = (0.0, 0.0, offset_monitor),
size = (td.inf, td.inf, 0),
name = 'far_field_monitor',
freqs = monitor_freq,
normal_dir = '+',
phi = np.linspace(0, 2*np.pi, 361),
theta = np.linspace(0, np.pi, 181)
)
Create the Simulation¶
sim = td.Simulation(
center = (0.0, 0.0, 0.0),
size = sim_size,
grid_spec = td.GridSpec.auto(
wavelength = (lambda_min + lambda_max)/2,
min_steps_per_wvl = automesh_per_wvl
),
run_time = t_stop,
sources = [dipole_source],
monitors = time_monitors + [far_field_monitor],
structures = [slab, etch],
medium = vacuum,
shutoff = 1e-7,
symmetry = [-1, 1, 0]
)
17:18:20 EDT WARNING: Structure: simulation.structures[0] (no `name` was specified) was detected as being less than half of a central wavelength from a PML on side x-min. To avoid inaccurate results or divergence, please increase gap between any structures and PML or fully extend structure through the pml.
WARNING: Suppressed 3 WARNING messages.
WARNING: Field projection monitor 'far_field_monitor' has observation points set up such that the monitor is projecting backwards with respect to its 'normal_dir'. If this was not intentional, please take a look at the documentation associated with this type of projection monitor to check how the observation point coordinate system is defined.
def gaussian_far_field_no_polarization_sin_debye_apod(theta, phi, NA, n=1.0):
"""
Compute an ideal far-field Gaussian beam profile with Debye apodization,
given the theta and phi angles, numerical aperture (NA), and refractive index (n).
Parameters:
phi, theta : 1D arrays (rad)
Input angular coordinates
NA, n : float
Numerical aperture and refractive index (of air)
Returns:
A : 2D array
Ideal far-field amplitude profile with Debye apodization
"""
theta_vals = np.asarray(theta)
phi_vals = np.asarray(phi)
theta_grid, phi_grid = np.meshgrid(theta_vals, phi_vals, indexing='ij')
# Gaussian term
gauss = np.exp(-(np.sin(theta_grid) / NA)**2)
# Debye apodization with safe clipping
apod = np.sqrt(np.clip(np.cos(theta_grid), 0, None))
return gauss * apod
def compute_gaussian_overlap_without_pol(
phi, theta, vals,
NA = 0.68,
n = 1.0,
theta_max_deg = None,
gauss_func = gaussian_far_field_no_polarization_sin_debye_apod,
):
"""
Compute the mode-overlap between the simulated far-field intensity (vals)
and an ideal Gaussian far-field WITHOUT polarization effects.
Overlap is computed as:
| ∫ E_sim E_gauss dΩ|^2 / ( ∫|E_sim|^2 dΩ ∫|E_gauss|^2 dΩ )
Added FEATURE:
If theta_max_deg is provided (in degrees), the integrals are evaluated
ONLY for theta <= theta_max_deg.
Parameters
----------
phi, theta : 1D arrays (rad)
Angular sampling coordinates of the simulated far-field.
vals : 2D array
Simulated intensity map, shape = (len(theta), len(phi)).
NA, n : floats
Gaussian beam NA and refractive index.
theta_max_deg : float or None
Upper limit of theta (in degrees) for restricting the overlap region.
If None → use entire theta-domain.
gauss_func : callable
Function that generates the ideal Gaussian amplitude:
gauss_func(theta, phi, NA, n) -> 2D array (len(theta), len(phi))
Returns
-------
percentage_overlap : float
Mode overlap (0–100%).
"""
# Convert inputs to arrays
phi = np.asarray(phi)
theta = np.asarray(theta)
vals = np.asarray(vals)
assert vals.shape == (len(theta), len(phi)), \
f"vals has shape {vals.shape}, but expected {(len(theta), len(phi))}"
# --- Choose which Gaussian model to use via gauss_func ---
E_gauss = gauss_func(theta, phi, NA, n)
I_gauss = E_gauss**2
# Sim amplitude = sqrt(intensity)
I_sim = vals
E_sim = np.sqrt(I_sim)
# Meshgrid for solid-angle element
theta_grid, phi_grid = np.meshgrid(theta, phi, indexing='ij')
# Solid angle element (assume uniform grid)
dtheta = float(np.mean(np.diff(theta)))
dphi = float(np.mean(np.diff(phi)))
dOmega = np.sin(theta_grid) * dtheta * dphi
# --------------------------------------
# Mask to restrict region if requested
# --------------------------------------
if theta_max_deg is not None:
theta_max_rad = np.radians(theta_max_deg)
mask = theta_grid <= theta_max_rad
else:
mask = np.ones_like(theta_grid, dtype=bool)
# Apply mask
E_sim_m = E_sim[mask]
E_gauss_m = E_gauss[mask]
I_sim_m = I_sim[mask]
I_gauss_m = I_gauss[mask]
dOmega_m = dOmega[mask]
# --------------------------
# Mode overlap numerator
# --------------------------
overlap = np.sum(E_sim_m * E_gauss_m * dOmega_m)
numerator = np.abs(overlap)**2
# --------------------------
# Normalization denominator
# --------------------------
norm_sim = np.sum(I_sim_m * dOmega_m)
norm_gauss = np.sum(I_gauss_m * dOmega_m)
denominator = norm_sim * norm_gauss
eta = numerator / denominator if denominator > 0 else 0.0
return float(100 * eta)
def compute_power_fraction_and_ratio(
phi, theta, vals,
NA_target,
NA_gauss=0.68,
n=1.0,
theta_max_deg=None,
gauss_func=gaussian_far_field_no_polarization_sin_debye_apod,
):
r"""
Compute the percentage of power captured within a target NA cone for:
(1) simulated far-field intensity vals(θ,φ)
(2) ideal Gaussian (amplitude) generated by gauss_func
Then compute the ratio:
ratio = (percent_sim / percent_gauss)
Power is integrated over solid angle:
P \propto \iint I(θ,φ) sinθ dθ dφ
Optional FEATURE:
If theta_max_deg is provided, both total-power integrals are computed
only over θ <= theta_max_deg (same as your overlap style).
Parameters
----------
phi, theta : 1D arrays (rad)
vals : 2D array
Simulated intensity map, shape = (len(theta), len(phi)).
NA_target : float
NA defining the capture cone for the fraction calculation.
NA_gauss : float
NA used to *define the Gaussian model* in gauss_func (beam divergence).
n : float
Refractive index of surrounding medium for θ = asin(NA/n).
theta_max_deg : float or None
If provided, restrict all integrals to θ <= theta_max_deg.
gauss_func : callable
Gaussian amplitude model:
gauss_func(theta, phi, NA_gauss, n) -> 2D amplitude array
Returns
-------
percent_sim : float
0–100 (%) power fraction within NA_target for simulated intensity.
percent_gauss : float
0–100 (%) power fraction within NA_target for Gaussian intensity |E|^2.
ratio : float
(percent_sim / percent_gauss); np.inf if percent_gauss == 0.
"""
if gauss_func is None:
raise ValueError("gauss_func must be provided (it should return Gaussian amplitude).")
# Convert inputs to arrays
phi = np.asarray(phi)
theta = np.asarray(theta)
vals = np.asarray(vals)
assert vals.shape == (len(theta), len(phi)), \
f"vals has shape {vals.shape}, but expected {(len(theta), len(phi))}"
# --- Build Gaussian intensity from amplitude ---
E_gauss = gauss_func(theta, phi, NA_gauss, n)
I_gauss = np.abs(E_gauss)**2
# Simulated intensity
I_sim = vals
# Meshgrid for solid-angle element (assume uniform grid like your function)
theta_grid, phi_grid = np.meshgrid(theta, phi, indexing='ij')
dtheta = float(np.mean(np.diff(theta)))
dphi = float(np.mean(np.diff(phi)))
dOmega = np.sin(theta_grid) * dtheta * dphi
# --------------------------------------
# Masks: (A) optional theta restriction
# --------------------------------------
if theta_max_deg is not None:
theta_max_rad = np.radians(theta_max_deg)
mask_theta = theta_grid <= theta_max_rad
else:
mask_theta = np.ones_like(theta_grid, dtype=bool)
# --------------------------------------
# Masks: (B) NA capture cone restriction
# --------------------------------------
theta_cap = np.arcsin(np.clip(NA_target / float(n), -1.0, 1.0))
mask_cap = theta_grid <= theta_cap
# Combined masks
mask_total = mask_theta
mask_in = mask_theta & mask_cap
# --------------------------
# Simulated power fractions
# --------------------------
P_sim_total = np.sum(I_sim[mask_total] * dOmega[mask_total])
P_sim_in = np.sum(I_sim[mask_in] * dOmega[mask_in])
frac_sim = (P_sim_in / P_sim_total) if P_sim_total > 0 else 0.0
# --------------------------
# Gaussian power fractions
# --------------------------
P_g_total = np.sum(I_gauss[mask_total] * dOmega[mask_total])
P_g_in = np.sum(I_gauss[mask_in] * dOmega[mask_in])
frac_gauss = (P_g_in / P_g_total) if P_g_total > 0 else 0.0
percent_sim = 100.0 * float(frac_sim)
percent_gauss = 100.0 * float(frac_gauss)
ratio = np.inf if percent_gauss == 0 else (percent_sim / percent_gauss)
return percent_sim, percent_gauss, float(ratio)
def make_far_field_plot_without_pol(phi, theta, vals, NA = 0.68, n=1.0,
gauss_func = gaussian_far_field_no_polarization_sin_debye_apod,
calculate_overlap=True):
"""
Plots the far-field intensity distribution, optionally saving the figure and calculating the mode overlap and mentioning it on the plot.
Parameters:
theta, phi : 1D arrays (rad)
Input angular coordinates
vals : 2D array
Far-field intensity values
NA, n : float
Numerical aperture and refractive index (of air)
calculate_overlap : bool
Whether to calculate and display the mode overlap (Ideally when plotting the Gaussian beam, this should be False)
"""
phi = np.asarray(phi)
theta = np.asarray(theta)
vals = np.asarray(vals)
assert vals.shape == (len(theta), len(phi))
if calculate_overlap:
overlap = compute_gaussian_overlap_without_pol(
phi, theta, vals,
NA=NA, n=n,
gauss_func=gauss_func
)
I = vals
I_norm = I / (I.max() if I.max()>0 else 1.0)
PHI, THETA = np.meshgrid(phi, theta, indexing = 'xy')
theta_NA_deg = np.degrees(np.arcsin(min(NA/float(n), 1.0)))
phi_circle = np.linspace(0, 2.0 * np.pi, 721)
theta_circle = np.full_like(phi_circle, theta_NA_deg)
fig1, ax1 = plt.subplots(subplot_kw={'projection': 'polar'}, figsize = (6.5,5.5))
ax1.tick_params(axis='y', colors='white')
pcm1 = ax1.pcolormesh(PHI, np.degrees(THETA), I_norm, shading='auto', cmap='magma')
cbar1 = fig1.colorbar(pcm1, ax=ax1, label=r"$|E|^2$")
ax1.set_ylim(0, 90)
#ax1.set_title('Far_field: Normalized Intensity')
ax1.plot(phi_circle, theta_circle, 'w--', lw = 2, label=f'NA={NA:.2f}, n = {n}')
ax1.legend(loc='upper right')
if calculate_overlap:
ax1.text(0.02, 0.98, f'Mode overlap: {overlap:.2f}%', transform = ax1.transAxes, va = 'top', ha = 'left',
bbox = dict(boxstyle = 'round, pad = 0.3', facecolor = 'black', alpha = 0.55), color = 'white')
fig1.tight_layout()
plt.show()
return
task_id = web.upload(sim, task_name = "Inverse_Designed_19th_Obj_6th_Stage_From_Epoch=5_Epoch=15_Rounded_UP_APW=40_T=30")
WARNING: Simulation has 2.36e+06 time steps. The 'run_time' may be unnecessarily large, unless there are very long-lived resonances.
17:18:21 EDT Created task 'Inverse_Designed_19th_Obj_6th_Stage_From_Epoch=5_Epoch=15_Rounded_ UP_APW=40_T=30' with resource_id 'fdve-054f38e1-2068-451e-b2a4-266ac88da4c4' and task_type 'FDTD'.
View task using web UI at 'https://tidy3d.simulation.cloud/workbench?taskId=fdve-054f38e1-206 8-451e-b2a4-266ac88da4c4'.
Task folder: 'default'.
Output()
17:18:23 EDT Estimated FlexCredit cost: 24.324. Minimum cost depends on task execution details. Use 'web.real_cost(task_id)' to get the billed FlexCredit cost after a simulation run.
Run the Simulation¶
web.start(task_id)
web.monitor(task_id, verbose = True)
17:18:24 EDT status = success
sim_data = web.load(task_id, verbose=False, path = "Inverse_Designed_19th_Obj_6th_Stage_From_Epoch=5_Epoch=15_Rounded_UP_APW=40_T=30.hdf5")
print(sim_data.log)
17:18:58 EDT WARNING: Structure: simulation.structures[0] (no `name` was specified) was detected as being less than half of a central wavelength from a PML on side x-min. To avoid inaccurate results or divergence, please increase gap between any structures and PML or fully extend structure through the pml.
WARNING: Suppressed 3 WARNING messages.
WARNING: Field projection monitor 'far_field_monitor' has observation points set up such that the monitor is projecting backwards with respect to its 'normal_dir'. If this was not intentional, please take a look at the documentation associated with this type of projection monitor to check how the observation point coordinate system is defined.
WARNING: Simulation final field decay value of 4.6e-07 is greater than the simulation shutoff threshold of 1e-07. Consider running the simulation again with a larger 'run_time' duration for more accurate results.
WARNING: Warning messages were found in the solver log. For more information, check 'SimulationData.log' or use 'web.download_log(task_id)'.
[18:45:02] WARNING: Structure: simulation.structures[0] (no `name` was
specified) was detected as being less than half of a central
wavelength from a PML on side x-min. To avoid inaccurate results or
divergence, please increase gap between any structures and PML or
fully extend structure through the pml.
WARNING: Suppressed 3 WARNING messages.
WARNING: Field projection monitor 'far_field_monitor' has observation
points set up such that the monitor is projecting backwards with
respect to its 'normal_dir'. If this was not intentional, please take
a look at the documentation associated with this type of projection
monitor to check how the observation point coordinate system is
defined.
USER: Simulation domain Nx, Ny, Nz: [800, 800, 134]
USER: Applied symmetries: (-1, 1, 0)
USER: Number of computational grid points: 2.1978e+07.
USER: Subpixel averaging method: SubpixelSpec(attrs={},
dielectric=PolarizedAveraging(attrs={}, type='PolarizedAveraging'),
metal=Staircasing(attrs={}, type='Staircasing'),
pec=PECConformal(attrs={}, type='PECConformal',
timestep_reduction=0.3, edge_singularity_correction=True),
pmc=Staircasing(attrs={}, type='Staircasing'),
lossy_metal=SurfaceImpedance(attrs={}, type='SurfaceImpedance',
timestep_reduction=0.0, edge_singularity_correction=True),
type='SubpixelSpec')
USER: Number of time steps: 2.3613e+06
USER: Automatic shutoff factor: 1.00e-07
USER: Time step (s): 1.2705e-17
USER:
USER: Compute source modes time (s): 0.1242
[18:45:03] USER: Rest of setup time (s): 0.9249
[18:45:31] USER: Compute monitor modes time (s): 0.0001
[20:15:58] USER: Solver time (s): 5450.9610
USER: Time-stepping speed (cells/s): 1.00e+10
[20:16:07] USER: Post-processing time (s): 8.4307
====== SOLVER LOG ======
Processing grid and structures...
Building FDTD update coefficients...
Solver setup time (s): 25.1779
Running solver for 2361336 time steps...
- Time step 435 / time 5.53e-15s ( 0 % done), field decay: 1.00e+00
- Time step 94453 / time 1.20e-12s ( 4 % done), field decay: 2.82e-03
- Time step 188906 / time 2.40e-12s ( 8 % done), field decay: 7.30e-05
- Time step 283360 / time 3.60e-12s ( 12 % done), field decay: 1.63e-03
- Time step 377813 / time 4.80e-12s ( 16 % done), field decay: 4.25e-04
- Time step 472267 / time 6.00e-12s ( 20 % done), field decay: 3.74e-04
- Time step 566720 / time 7.20e-12s ( 24 % done), field decay: 7.02e-04
- Time step 661174 / time 8.40e-12s ( 28 % done), field decay: 8.19e-06
- Time step 755627 / time 9.60e-12s ( 32 % done), field decay: 3.99e-04
- Time step 850080 / time 1.08e-11s ( 36 % done), field decay: 1.34e-04
- Time step 944534 / time 1.20e-11s ( 40 % done), field decay: 7.57e-05
- Time step 1038987 / time 1.32e-11s ( 44 % done), field decay: 1.88e-04
- Time step 1133441 / time 1.44e-11s ( 48 % done), field decay: 3.73e-06
- Time step 1227894 / time 1.56e-11s ( 52 % done), field decay: 9.55e-05
- Time step 1322348 / time 1.68e-11s ( 56 % done), field decay: 4.35e-05
- Time step 1416801 / time 1.80e-11s ( 60 % done), field decay: 1.47e-05
- Time step 1511255 / time 1.92e-11s ( 64 % done), field decay: 4.99e-05
- Time step 1605708 / time 2.04e-11s ( 68 % done), field decay: 1.88e-06
- Time step 1700161 / time 2.16e-11s ( 72 % done), field decay: 2.25e-05
- Time step 1794615 / time 2.28e-11s ( 76 % done), field decay: 1.29e-05
- Time step 1889068 / time 2.40e-11s ( 80 % done), field decay: 2.70e-06
- Time step 1983522 / time 2.52e-11s ( 84 % done), field decay: 1.29e-05
- Time step 2077975 / time 2.64e-11s ( 88 % done), field decay: 8.37e-07
- Time step 2172429 / time 2.76e-11s ( 92 % done), field decay: 5.02e-06
- Time step 2266882 / time 2.88e-11s ( 96 % done), field decay: 3.75e-06
- Time step 2361335 / time 3.00e-11s (100 % done), field decay: 4.60e-07
Time-stepping time (s): 5186.8286
Field projection time (s): 232.2404
Data write time (s): 6.7131
sim_data = td.SimulationData.from_file(fname = "Inverse_Designed_19th_Obj_6th_Stage_From_Epoch=5_Epoch=15_Rounded_UP_APW=40_T=30.hdf5")
17:18:59 EDT WARNING: Structure: simulation.structures[0] (no `name` was specified) was detected as being less than half of a central wavelength from a PML on side x-min. To avoid inaccurate results or divergence, please increase gap between any structures and PML or fully extend structure through the pml.
WARNING: Suppressed 3 WARNING messages.
WARNING: Field projection monitor 'far_field_monitor' has observation points set up such that the monitor is projecting backwards with respect to its 'normal_dir'. If this was not intentional, please take a look at the documentation associated with this type of projection monitor to check how the observation point coordinate system is defined.
Analyze Results¶
fig, ax = plt.subplots(1, 1, tight_layout = True, figsize = (6,4))
time_monitor_names = [monitor.name for monitor in time_monitors]
time_response_Ex = np.zeros_like(sim_data["time_monitor_1"].Ex.squeeze())
for monitor_name in time_monitor_names:
time_response_Ex += sim_data[monitor_name].Ex.squeeze()
time_response_Ex /= len(time_monitor_names)
freq_response_Ex = np.abs(np.fft.fft(time_response_Ex))
freqs_Ex = np.linspace(0, 1/sim_data.simulation.dt, len(time_response_Ex))
wavelengths_Ex = td.constants.C_0 / freqs_Ex
plot_inds_Ex = np.where((monitor_freq[-1] < freqs_Ex) & (freqs_Ex < monitor_freq[0]))
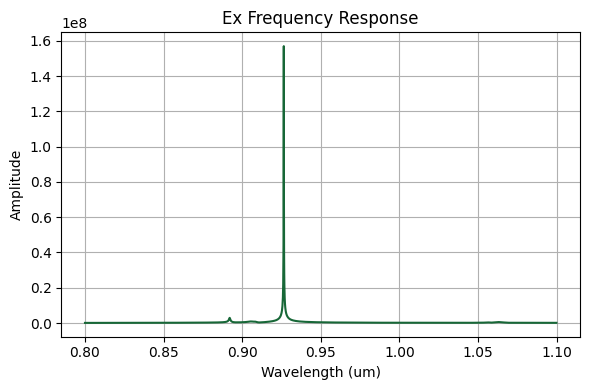
ax.plot(wavelengths_Ex[plot_inds_Ex], freq_response_Ex[plot_inds_Ex])
ax.set_xlabel("Wavelength (um)")
ax.set_ylabel("Amplitude")
ax.set_title("Ex Frequency Response")
ax.grid(True)
plt.show()
/var/folders/qn/syhrzy8n7930sgqvxv2x65s40000gn/T/ipykernel_59250/3831151282.py:15: RuntimeWarning: divide by zero encountered in divide wavelengths_Ex = td.constants.C_0 / freqs_Ex
resonance_finder = ResonanceFinder(freq_window = (monitor_freq[-1], monitor_freq[0]))
resonance_data = resonance_finder.run(signals = sim_data.data)
df_actual = resonance_data.to_dataframe().reset_index()
df_actual['wavelength'] = td.constants.C_0 / df_actual['freq']
print(df_actual[['freq', 'wavelength', 'decay', 'Q', 'amplitude', 'phase', 'error']])
freq wavelength decay Q amplitude phase \
0 2.699416e+14 1.110583 2.142817e+12 395.762551 30.476848 2.108739
1 2.746487e+14 1.091549 2.879579e+12 299.638982 41.040942 -2.805304
2 2.833892e+14 1.057883 1.830160e+12 486.456684 12.831113 -3.017307
3 3.235854e+14 0.926471 1.137087e+11 8940.158494 254.820183 -0.476974
4 3.298842e+14 0.908781 3.083029e+12 336.150431 88.478080 -0.707611
5 3.309785e+14 0.905776 3.699974e+12 281.028907 99.558649 0.086578
6 3.360605e+14 0.892079 9.866268e+11 1070.075330 160.566584 0.696290
7 3.525750e+14 0.850294 2.285641e+12 484.611064 8.606144 1.830012
8 4.125453e+14 0.726690 1.534290e+12 844.722658 418.070821 -2.465529
error
0 0.004749
1 0.006332
2 0.006637
3 0.000090
4 0.004032
5 0.005352
6 0.001005
7 0.007987
8 0.002071
indices = np.where((monitor_lambda >= 0.92) & (monitor_lambda <=0.93))[0]
for idx in indices:
print(f"Index: {idx}, Wavelength: {monitor_lambda[idx]:.6f} um")
Index: 160, Wavelength: 0.920000 um Index: 161, Wavelength: 0.920750 um Index: 162, Wavelength: 0.921500 um Index: 163, Wavelength: 0.922250 um Index: 164, Wavelength: 0.923000 um Index: 165, Wavelength: 0.923750 um Index: 166, Wavelength: 0.924500 um Index: 167, Wavelength: 0.925250 um Index: 168, Wavelength: 0.926000 um Index: 169, Wavelength: 0.926750 um Index: 170, Wavelength: 0.927500 um Index: 171, Wavelength: 0.928250 um Index: 172, Wavelength: 0.929000 um Index: 173, Wavelength: 0.929750 um
Far-Field Analysis¶
f_index = 168
far_field_data = sim_data[far_field_monitor.name]
Etheta = far_field_data.Etheta.isel(f = f_index, r=0).values
Ephi = far_field_data.Ephi.isel(f = f_index, r=0).values
phi_vals = far_field_data.phi
theta_vals = far_field_data.theta
intensity = np.abs(Etheta)**2 + np.abs(Ephi)**2
compute_gaussian_overlap_without_pol(
phi_vals, theta_vals, intensity,
NA = 0.68, n = 1.0,
gauss_func=gaussian_far_field_no_polarization_sin_debye_apod
)
86.88443254695612
p_sim, p_g, r = compute_power_fraction_and_ratio(
phi_vals, theta_vals, vals=intensity,
NA_target=0.68,
NA_gauss=0.68,
n=1.0,
theta_max_deg=90,
gauss_func=gaussian_far_field_no_polarization_sin_debye_apod
)
print(p_sim, p_g, r)
76.16387279212572 87.2671390916859 0.8727669267592844
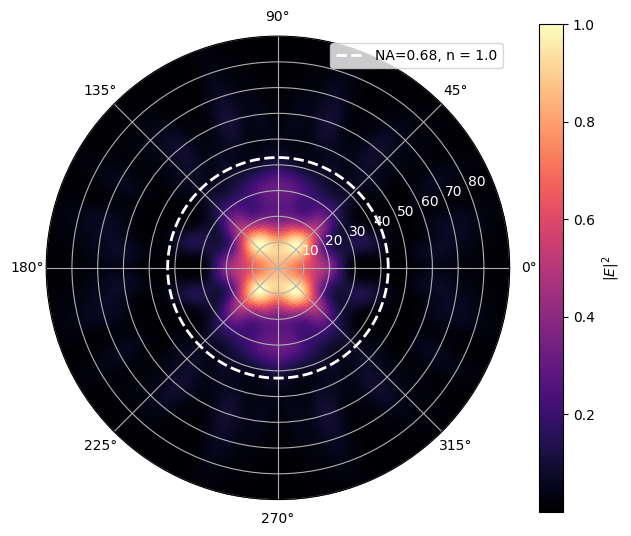
make_far_field_plot_without_pol(
phi_vals, theta_vals, intensity,
NA = 0.68, n=1.0,
gauss_func=gaussian_far_field_no_polarization_sin_debye_apod,
calculate_overlap=False
)