Author: Leyang Liu, University of Illinois Urbana-Champaign
In this notebook, we simulate the optical reflection spectrum of a 1D photonic crystal / grating structure using Tidy3D. The structure consists of patterned SiO₂ and TiO₂ layers embedded in polymer (PVA) on a substrate. A broadband plane wave is incident on the structure, and the notebook computes:
- Reflection spectrum vs wavelength
- Electric field distribution
- Poynting vector flow
This type of simulation is typical for biosensors, dielectric metasurfaces, or photonic crystal reflectors.

import matplotlib.pyplot as plt
import numpy as np
import tidy3d as td
import tidy3d.web as web
from tidy3d.plugins.dispersion import FastDispersionFitter
15:53:46 UTC WARNING: Using canonical configuration directory at '/home/tidy3d/.config/tidy3d'. Found legacy directory at '~/.tidy3d', which will be ignored. Remove it manually or run 'tidy3d config migrate --delete-legacy' to clean up.
Define material properties:
# materials
SiO2 = td.Medium(
name = 'SiO2',
permittivity = 2.1025,
)
TiO2 = td.Medium(
name = 'TiO2',
permittivity = 5.8081000000000005,
)
background = td.Medium(permittivity=1**2)
PVA = td.Medium(permittivity = 1.483**2)
Define structure geometry:
period = 0.39
TiO2_thick = 0.105
SiO2_thick = 0.1
PVA_thick = 0.070
PVA_bottom_thick = 0.048
sub_thick = 2
monitor_distance = 1.0
monitor_gap = 0.1
dutycycle_SiO2 = 0.55
TiO2_bottom_thick = 0.067
slope_angle = 70/180*np.pi
dutycycle_TiO2 = 0.6
size_z = sub_thick+SiO2_thick+TiO2_thick+monitor_distance+monitor_gap
SiO2_top_width = dutycycle_SiO2*period
SiO2_bottom_width = SiO2_top_width+2*SiO2_thick/np.tan(slope_angle)
TiO2_top_width = dutycycle_TiO2*period
TiO2_bottom_width = TiO2_top_width+2*TiO2_thick/np.tan(slope_angle)
Define simulation parameters:
# the epoxy layer top surface is at z=0
sim_center = (0, 0, 0)
sim_size = (
period,
0,
size_z
)
# wavelength / frequency setup
nm = 1e-3
wavelength_min = 500 * nm
wavelength_max = 700 * nm
freq_min = td.C_0 / wavelength_max
freq_max = td.C_0 / wavelength_min
freq0 = td.C_0 / (500 * nm)
fwidth = freq_max - freq_min
run_time = 10e-11
# SiO2 sub
SiO2_sub = td.Structure(
geometry=td.Box(
center=[0.0, 0.0, -0.5 * sub_thick],
size=[td.inf, td.inf, sub_thick],
),
medium=SiO2,
name="SiO2_sub",
)
# SiO2 teeth
vertices1 = [[SiO2_bottom_width / 2, 0],
[-SiO2_bottom_width / 2, 0],
[-SiO2_top_width / 2, SiO2_thick],
[SiO2_top_width / 2, SiO2_thick]]
SiO2_teeth = td.Structure(
geometry = td.PolySlab(axis = 1, slab_bounds = [-td.inf, td.inf], vertices = vertices1),
name = 'SiO2_teeth',
medium = SiO2
)
# TiO2
vertices2 = [[TiO2_bottom_width / 2, TiO2_bottom_thick],
[TiO2_top_width / 2, SiO2_thick + TiO2_thick],
[-TiO2_top_width / 2, SiO2_thick + TiO2_thick],
[-TiO2_bottom_width / 2, TiO2_bottom_thick],
[-period / 2, TiO2_bottom_thick],
[-period / 2, 0],
[-SiO2_bottom_width / 2, 0],
[-SiO2_top_width / 2, SiO2_thick],
[SiO2_top_width / 2, SiO2_thick],
[SiO2_bottom_width / 2, 0],
[period / 2, 0],
[period / 2, TiO2_bottom_thick]]
TiO2 = td.Structure(
geometry = td.PolySlab(axis = 1, slab_bounds = [-td.inf, td.inf], vertices = vertices2),
medium = TiO2,
name = 'TiO2'
)
# PVA
vertices3 = [[TiO2_bottom_width / 2, TiO2_bottom_thick + PVA_bottom_thick],
[TiO2_top_width / 2, SiO2_thick + TiO2_thick + PVA_thick],
[-TiO2_top_width / 2, SiO2_thick + TiO2_thick + PVA_thick],
[-TiO2_bottom_width / 2, TiO2_bottom_thick + PVA_bottom_thick],
[-period / 2, TiO2_bottom_thick + PVA_bottom_thick],
[-period / 2, TiO2_bottom_thick],
[-TiO2_bottom_width / 2, TiO2_bottom_thick],
[-TiO2_top_width / 2, SiO2_thick + TiO2_thick],
[TiO2_top_width / 2, SiO2_thick + TiO2_thick],
[TiO2_bottom_width / 2, TiO2_bottom_thick],
[period / 2, TiO2_bottom_thick],
[period / 2, TiO2_bottom_thick + PVA_bottom_thick]]
PVA = td.Structure(
geometry = td.PolySlab(axis = 1, slab_bounds = [-td.inf, td.inf], vertices = vertices3),
medium = PVA,
name = 'PVA'
)
# the order here matters, because the teeth must override the bottom film layer
geometry = [SiO2_sub, SiO2_teeth, TiO2, PVA]
# geometry = [epoxy_layer, bottom_film, grating_teeth, top_film]
# boundary conditions: the simulation is periodic in the x-y plane, and simulates
# an infinite domain along z
boundary_spec = td.BoundarySpec(
x=td.Boundary.periodic(),
y=td.Boundary.periodic(),
z=td.Boundary.pml(),
)
# grid specification
grid_spec = td.GridSpec.auto(min_steps_per_wvl=30)
Define source:
source_time = td.GaussianPulse(freq0=freq0, fwidth=fwidth)
source = td.PlaneWave(
center=[0, 0, TiO2_thick + SiO2_thick + monitor_distance],
size=[td.inf, td.inf, 0.0],
source_time=source_time,
pol_angle=0,
# pol_angle=np.pi/2,
direction="-",
angle_theta=np.deg2rad(0),
angular_spec=td.FixedAngleSpec(),
)
Define monitors:
# create field monitor
monitor_xz = td.FieldMonitor(
center=sim_center,
size=[td.inf, 0, td.inf],
freqs=[freq0],
name="fields_xz",
)
# create flux monitors
freqs = np.linspace(freq_min, freq_max, 1000)
monitor_flux_refl = td.FluxMonitor(
center=[0, 0, monitor_distance],
size=[td.inf, td.inf, 0.0],
freqs=freqs,
name="flux_refl",
)
monitors = [monitor_xz, monitor_flux_refl]
Put everything together in a simulation object:
# create the simulation
sim = td.Simulation(
center=sim_center,
size=sim_size,
grid_spec=grid_spec,
structures=geometry,
sources=[source],
monitors=monitors,
run_time=run_time,
boundary_spec=boundary_spec,
medium=background,
shutoff=1e-7,
)
# plot the simulation domain
#sim.plot_3d()
#plt.show()
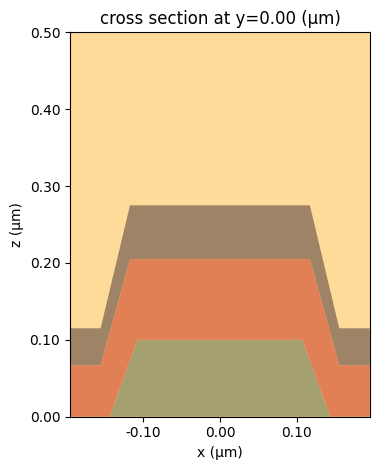
sim.plot(y=0.0, vlim=[0, 0.5])
plt.show()
15:53:49 UTC WARNING: Structure at 'structures[2]' has bounds that extend exactly to simulation edges. This can cause unexpected behavior. If intending to extend the structure to infinity along one dimension, use td.inf as a size variable instead to make this explicit.
WARNING: Suppressed 3 WARNING messages.
import tidy3d.web as web
sim_data = web.run(sim, task_name="biosensor", path="data/biosensor.hdf5", verbose=True)
WARNING: Simulation has 6.33e+06 time steps. The 'run_time' may be unnecessarily large, unless there are very long-lived resonances.
Created task 'biosensor' with resource_id 'fdve-9878e97e-7c50-4fd4-8631-65f84387e98b' and task_type 'FDTD'.
View task using web UI at 'https://tidy3d.simulation.cloud/workbench?taskId=fdve-9878e97e-7c5 0-4fd4-8631-65f84387e98b'.
Task folder: 'default'.
Output()
15:53:50 UTC Estimated FlexCredit cost: 0.040. Minimum cost depends on task execution details. Use 'web.real_cost(task_id)' to get the billed FlexCredit cost after a simulation run.
status = success
Output()
15:53:51 UTC Loading simulation from data/biosensor.hdf5
WARNING: Structure at 'structures[2]' has bounds that extend exactly to simulation edges. This can cause unexpected behavior. If intending to extend the structure to infinity along one dimension, use td.inf as a size variable instead to make this explicit.
WARNING: Suppressed 3 WARNING messages.
WARNING: Warning messages were found in the solver log. For more information, check 'SimulationData.log' or use 'web.download_log(task_id)'.
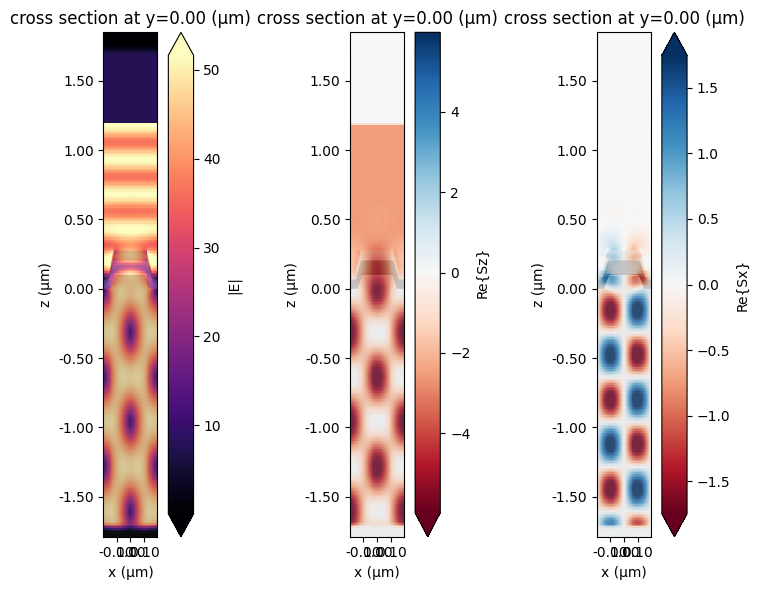
fig, ax = plt.subplots(1, 3, figsize=(8, 6), tight_layout=True)
sim_data.plot_field("fields_xz", field_name="E", val="abs", f=freq0, ax=ax[0])
sim_data.plot_field("fields_xz", field_name="Sz", val="real", f=freq0, ax=ax[1])
sim_data.plot_field("fields_xz", field_name="Sx", val="real", f=freq0, ax=ax[2])
plt.show()
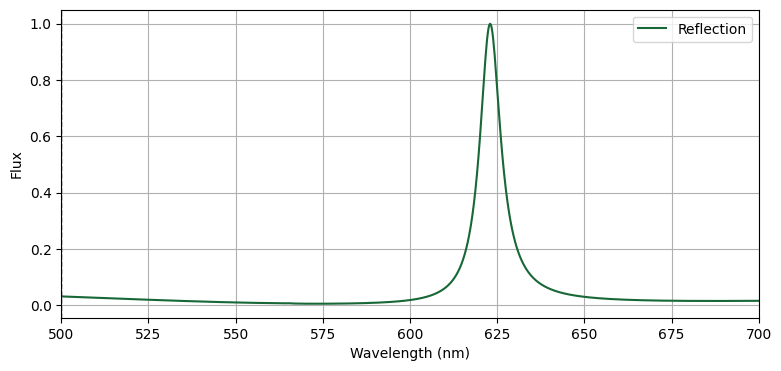
reflection = sim_data["flux_refl"].flux+1
fig, ax = plt.subplots(figsize=(9, 4))
ax.plot(td.C_0 / freqs * 1e3, reflection, label="Reflection")
# wavelength for the field plots
ax.axvline(td.C_0 / freq0 * 1e3, ls="--", color="k", lw=1)
ax.set(
xlabel="Wavelength (nm)",
ylabel="Flux",
xlim=(wavelength_min * 1e3, wavelength_max * 1e3),
)
ax.legend()
ax.grid()
#plt.xlim(550, 650)
plt.show()
#fig.savefig("1degree_30.tif", dpi=300)